COVID-19 research goes public through new portal
 (Download Image)
(Download Image)
A new online data portal is making available to the public a wealth of data Lawrence Livermore National Laboratory scientists have gathered from their ongoing COVID-19 molecular design projects, particularly the computer-based “virtual” screening of small molecules and designed antibodies for interactions with the SARS-CoV-2 virus for drug design purposes.
To help accelerate discovery of therapeutic antibodies or antiviral drugs for SARS-CoV-2, the virus that causes COVID-19, Lawrence Livermore National Laboratory (LLNL) has launched a searchable data portal to share its COVID-19 research with scientists worldwide and the general public.
The portal houses a wealth of data LLNL scientists have gathered from their ongoing COVID-19 molecular design projects, particularly the computer-based “virtual” screening of small molecules and designed antibodies for interactions with the SARS-CoV-2 virus for drug design purposes. The data is queryable by criteria such as chemical structure and binding probability scores, so outside researchers can easily locate relevant data for their own work.
“We do hope that somebody can use our information to help them make a small molecule drug or a better antibody,” said LLNL scientist Felice Lightstone, who is leading the anti-viral research. “We took a lot of time and effort to make these predictions, and we want to share them with the world so other researchers don’t have to redo the work and can advance their own research. It’s all about speeding up the process.”
Specifically, the initial data release includes predicted antibody sequences and homology models of viral proteins from the viral genome, small molecule compounds from public sources, viral protein targets and co-complexes of small molecule binding to viral proteins with predicted scores from binding energy and machine learning models. Predictions of drug safety and pharmacokinetics (how compounds distribute in the body) will be released in version two of the portal. Most results can be queried by chemical compound or protein and can be filtered by functional group or prediction scores. The portal is based on a system developed through a collaboration with the American Heart Association.
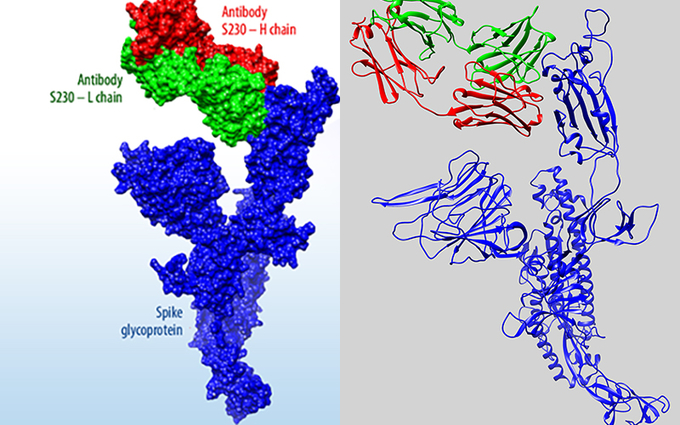
“With our released template structures, our calculations can be used to inform investigations of other antibodies targeting the same regions on the viral spike protein or antibodies derived from the same template antibodies we used,” said staff scientist Tom Desautels, principal investigator for the antibody project. “These results are also useful to researchers interested in how to integrate machine learning and computational structural biology. This sort of community science and feedback is really beneficial to us too, because it helps us to refine our process and methods.”
The portal will be regularly updated and will, in a few months, provide the results of experiments performed at the Laboratory on the effectiveness of small molecules and antibodies against SARS-CoV-2.
The data portal team includes Marisa Torres, Jonathan Allen, Dan Kirshner, Aidan Epstein, Masha Aseeva and the Green Data Oasis Team from Livermore Computing.
Contact
 Jeremy Thomas
Jeremy Thomas
[email protected]
(925) 422-5539
Related Links
LLNL’s COVID-19 researchTags
Bioscience and BioengineeringBiosciences and Biotechnology
HPC, Simulation, and Data Science
Computing
Physical and Life Sciences
Featured Articles







