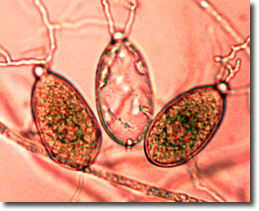
DOE Joint Genome Institute decodes DNA of two plant-killing pathogens
WALNUT CREEK, Calif. — By comparing the complete genome sequences of two plant-killing pathogens and related organisms, researchers from the U.S. Department of Energy Joint Genome Institute (DOE JGI), in collaboration with the Virginia Bioinformatics Institute (VBI) and others, have uncovered crucial aspects of the disease-causing mechanisms of “sudden oak death” (SOD) and soybean root rot disease.
The research, the result of a four-year, $4 million multi-agency project supported by DOE, U.S. Department of Agriculture (USDA), and the National Science Foundation (NSF), appears in the Sept. 1, 2006, edition of the journal Science .
“This project best exemplifies how the capabilities that were established at the DOE JGI for sequencing the human genome are now proving to be essential for addressing important environmental challenges,” said Eddy Rubin, DOE JGI Director. “We are now capable of rapidly responding to the urgent needs of the nation’s largest industry, agriculture, where genome sequence information can be brought to bear on characterizing such economically important microorganisms as those that cause sudden oak death and soybean root rot. For these pathogens, the genome sequence is the wiring diagram of the cellular processes that can be targeted for novel detection systems and for safe and effective means of control.”
Phytophthora (pronounced “Fy-TOFF-thor-uh”) species, the target pathogens, attack a wide variety of plants, including agricultural crops as well as trees and shrubs of native ecosystems and backyard gardens alike. Phytophthora ramorum (“ruh-MORE-um”) causes Sudden Oak Death, and Phytophthora sojae (“SEW-jay”) attacks primarily soybeans.
“Among the discoveries embedded in the DNA of Phytophthora ramorum is the presence of more than 13,000 diagnostically different single-letter changes that vary among strains of the disease,” said Jeffrey Boore, senior author of the Science study and DOE JGI Evolutionary Genomics Program head. “These ‘fingerprints’ are already being used to track the movement and identity of these species as they progressively invade different regions. Knowing the pattern of the pathogen’s spread may help researchers to design strategies for thwarting it.
“Another great benefit that has come from access to the genome sequence has been the design and production of technologies – now-available gene chips – that allow us to monitor the expression patterns of thousands of genes simultaneously as these change during the course of infection and other life stages of these organisms,” Boore said. “Together, these advances promise to reveal processes that can be blocked by human intervention and thus prevent the devastating economic losses to soybean crops and the environmental destruction of woodlands.”
The researchers observed that the pathogens, during the initial hours of infection, derive their nutrition from the living plant tissue, but after the infection has been established, they switch to fueling their growth from the killed plant tissue. They hypothesized that the two species produce gene products that enable them to evade or suppress the plant’s defense responses during infection and later produce gene products that kill and destroy plant tissue.
“What is extraordinary about the Phytophthora genomes is that almost half of the genes contained in them show signs of rapid adaptation,” said VBI Professor Brett Tyler, the project’s coordinator and lead author of the Science paper. “We speculate that the rapidly changing genes are being driven to evolve by pressure from the defense systems of the pathogens’ host plants,” Tyler said. “The unprecedented level of genetic flexibility in these organisms gives us insights into how these pathogens have become successful. At the same time it has helped us identify weak points in the organisms that can be targeted to control them.”
“These results, which identify the DNA sequence of the two pathogens, arm scientists and practitioners to assist in the development of new Phytophthora disease-control measures,” said Dr. Gale Buchanan, USDA Undersecretary for Research, Education and Economics. “Defeating these two pathogens could significantly reduce the billions of dollars lost to crop damage worldwide each year.”
Sudden oak death was first reported in 1995; The agent responsible for the disease was discovered by University of California scientists in 2000. The pathogen is known to be present in more than a dozen California counties and also in southern Oregon. It has also been detected at scores of nurseries across the nation, elevating concerns about the pathogen to an all-time high.
Symptoms vary depending on the host. Infected oak trees exhibit oozing cankers on the trunk, and often succumb to the disease or to secondary infections as they are weakened by P. ramorum . In leaves, the pathogen reveals its presence through blight and twig dieback. In this manifestation, the disease can be transmitted by such plants as California bay laurel, camellia, and rhododendron.
The economic impact of Phytophthora sojae is far-reaching. The U.S. produces almost half the world’s soybeans. Losses attributed to P. sojae infestation, soybean root rot, exceed $1 billion annually. Soybean, the world’s most valuable legume crop, is of particular interest to DOE because it is the principal source of biodiesel, a renewable, alternative fuel. Biodiesel has the highest energy content of any alternative fuel and is significantly more environmentally friendly than comparable petroleum-based fuels, since it degrades rapidly in the environment. Earlier this year, DOE and USDA announced that they will share resources and coordinate the study of plant and microbial genomics, and as a result DOE JGI will tackle the sequencing of the soybean genome as the first project.
“The Phytophthoras , in addition to their great economic importance, are fascinating organisms with very distinct and interesting biology,” said Maryanna Henkart, NSF’s division director for Molecular and Cellular Biosciences. “These new genome sequences will contribute to our basic understanding of normal plant-microbe relationships as well as their roles in disease. Generating such fundamental knowledge is at the core of NSF's mission, and we are pleased to have played a role in promoting this important project,” Henkart said.
DOE JGI, using the iterative whole-genome shotgun approach, generated nine-fold coverage of the 95 million nucleic acid bases, or units of the genetic code, of the P. sojae genome and seven-fold coverage of the 65-million-base P. ramorum genome.
The aptly named genus Phytophthora derives its moniker from the Greek words for “plant destroyer.” Part of a fungus-like group of organisms known as oomycetes , or water molds, they are relatives of such aquatic algae as diatoms and kelp. The pathogens survive as thick-walled spores that can persist in soil for years. Of the 59 recognized Phytophthora species, it was P. infestans that was responsible for the mid-19th century Irish potato famine.
The Virginia Bioinformatics Institute (VBI) at Virginia Tech has a research platform centered on understanding the “disease triangle” of host-pathogen-environment interactions in plants, humans and other animals. By successfully channeling innovation into transdisciplinary approaches that combine information technology and biology, researchers at VBI are addressing some of today’s key challenges in the biomedical, environmental and plant sciences.
The DOE Joint Genome Institute, supported by the DOE Office of Science, unites the expertise of five national laboratories – Lawrence Livermore, Lawrence Berkeley, Los Alamos, Oak Ridge, and Pacific Northwest – along with the Stanford Human Genome Center to advance genomics in support of the DOE mission related to clean energy generation and environmental characterization and clean-up. DOE JGI’s Walnut Creek, Calif., Production Genomics Facility provides integrated high-throughput sequencing and computational analysis that enable systems-based scientific approaches to these challenges. Click here for additional information about DOE JGI.
Contact
David Gilbert[email protected]
925-296-5643
Related Links
“PhytophthoraGenome Sequences Uncover Evolutionary Origins and Mechanisms of Pathogenesis”“Genomes Highlight Plant Pathogens’ Powerful Arsenal”
“Genome Info from ‘Plant Destroyers’ Could Save Trees, Beans and Chocolate”
DOE JGI to decode DNA of soybean,a key alternative energy source