Overlapping genes for prolonged genetic circuit stability
 (Download Image)
(Download Image)
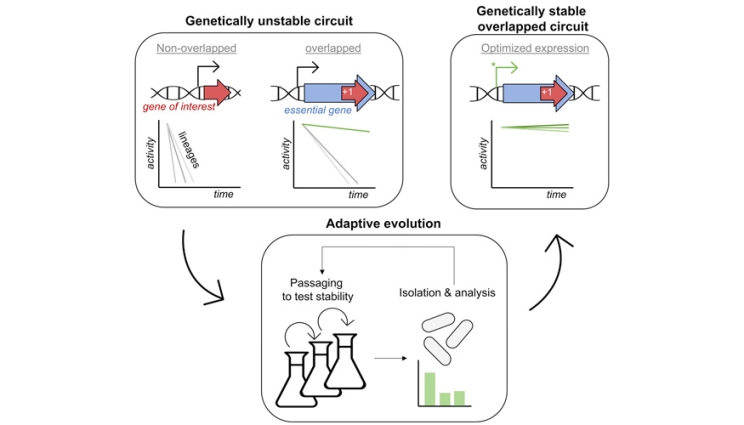
Graphical illustration comparing a non-overlapped gene circuit with an overlapped one and showing how, once overlapped, gene stability is optimized.
For the past several decades, synthetic biologists have sought to genetically engineer microorganisms for a wide range of application—including therapeutics discovery and delivery, drug manufacturing, agricultural yields, biofuel production, mineral extraction, and waste degradation. This is achieved through the design of genetic circuits, which are made up of DNA parts that enable a cell to perform a desired function. For example, microbes can be genetically engineered to produce proteins that enhance nutrient acquisition or drought resistance in agricultural crops.
Such biotechnology applications require the stable function of genetic circuits over extended time periods to allow for robust production of a desired product. Oftentimes, engineered genetic circuits can adversely affect the fitness of the host cell, and as a result, prematurely fail due to inactivating mutations. Therefore, the development of tools that mitigate genetic instability while maintaining circuit function is necessary and central to synthetic biology.
Recently, approaches using gene overlaps have been developed to enhance genetic stability, where a gene of interest (GOI) and an essential gene (critical genes for organism survival) are synthetically encoded (“entangled”) within the same DNA sequence to constrain the evolutionary outcomes of the GOI. To this end, a team of scientists conducted a study, published in Nucleic Acids Research, demonstrating the use of gene entanglement for improving genetic circuit stability, particularly for burdensome components that are prone to mutational inactivation.
The team’s findings provide insights into how gene entanglement alters the allowable mutation possibilities and the evolutionary trajectory of synthetic circuits. By aligning the function of the genetic circuit directly with cell survival, mutations that would otherwise disrupt circuit function were disallowed. Interestingly, this entanglement approach yielded mutations that reduced the fitness burden on the host cell without overriding circuit function. The result was a genetic circuit that remained stable over 20 consecutive days of growth. Altogether, these results showcase how sequence entanglement can be used to harness evolution for genetic circuit optimization.
[J.L Chlebek, S.P Leonard, C. Kang-Yun, M.C Yung, D.P Ricci, Y. Jiao, D.M. Park, Prolonging Genetic Circuit Stability Through Adaptive Evolution of Overlapping Genes, Nucleic Acids Research (2023), DOI: 10.1093/nar/gkad484.]
–Physical and Life Sciences Communications Team
Tags
Bioscience and BioengineeringBiosciences and Biotechnology
Physical and Life Sciences
Featured Articles







