Multi-lab research to improve COVID-19 diagnostics
 (Download Image)
(Download Image)
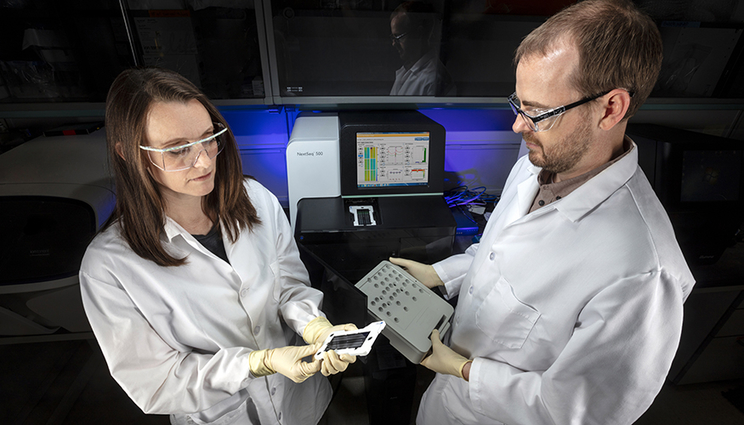
LLNL biologists Jessica Wollard and James Thissen use molecular-based technologies, such as PCR, microarray and genomic sequencing, to characterize microbes and pathogens in samples.
In response to the ongoing need for COVID-19 testing, Lawrence Livermore National Laboratory (LLNL) biologists are part of a collaborative research effort focused on improving the speed and accuracy of diagnostic tests, while enhancing the ability to adapt diagnostic tools as the virus evolves.
Currently, the fastest way to identify known pathogens is by using a DNA-based detection technology known as polymerase chain reaction (PCR), which can identify pathogens in biological samples in a matter of minutes to hours. The technology’s high sensitivity enables virus detection soon after infection, even before the onset of disease.
However, viruses such as SARS-CoV-2 do not contain DNA. They only contain RNA. Thus, once the RNA is extracted from a respiratory sample obtained via nasal or oral swabs, a reverse transcription PCR technique is used to reverse-transcribe the RNA into DNA. Then PCR technology is used to identify the virus by its genetic code.
Due to the increased demand for COVID-19 tests that use PCR technology, there is a critical shortage of commercially available RNA extraction kits that are approved under an Emergency Use Authorization (EUA) by the Food and Drug Administration (FDA). LLNL scientists are exploring alternatives to RNA extraction methods to address this shortage.
LLNL biologist Crystal Jaing is leading several aspects of the multi-lab research project, including efforts to identify and validate alternate RNA extraction techniques that do not rely on commercially available kits, thereby providing an alternative solution to the reagent shortage. For example, they will explore direct conversion of clinical samples into the form needed to analyze them using PCR, without the need for extraction.
In addition, the research team, which includes LLNL biologists Jessica Wollard, James Thissen and Aubree Hinckley, along with scientists from seven other DOE laboratories, will validate COVID-19 detection metrics on alternate diagnostic instrumentation — those instruments that are not already approved under a FDA EUA. They also will identify expanded protocols that can be used by clinical laboratories for COVID-19 testing. For example, the protocols will address issues such as optimal procedures for sample handling and purification for various types of existing PCR instrumentation used at clinical laboratories and public health departments.
Screening for co-infections
The research team also is studying how the presence of other viruses, bacteria and fungal pathogens affect how someone responds when infected with the SARS-CoV-2 virus. “We anticipate being able to identify microbes that may impact a person’s susceptibility to COVID-19 infection, or promote resilience,” Jaing said. The study will provide insight regarding how to diagnose and treat individuals infected by SARS-CoV-2, especially as the virus evolves and genomic changes occur.
Scientists will use an existing microbial detection technology developed at LLNL to analyze COVID-19 oral and nasal samples and identify the presence of any other pathogens. The Lawrence Livermore Microbial Detection Array is a high-throughput, nucleic-acid detection technology that can analyze 12,000 microbial species in a single test. In addition, LLNL scientists will work with other DOE laboratories to conduct metagenomic sequencing and use the Livermore Metagenomics Analysis Toolkit to confirm the presence of pathogens.
LLNL biostatistician Nisha Mulakken is part of a team that is building powerful analytical tools to evaluate diagnostic testing methods as the virus evolves. For example, she will mine data regarding emerging hotspots around the world using tools such as nextstrain.org.
Creating a knowledge base of diagnostic testing
Jaing’s role in the DOE-funded project also includes efforts to develop an integrated data science tool that will benefit the COVID-19 research community. LLNL biostatistician Nisha Mulakken and bioinformaticist Marisa Torres are part of the team developing the data repository, which will include information regarding the microbes present in clinical samples obtained from COVID-19 patients, as well as genomic data regarding virus mutations.
“Our plan is to create a knowledge base of genomic models to facilitate diagnostic assay standardization,” Jaing said. “We hope this unique resource will help investigators answer key questions such as which diagnostic sequences are most effective at identifying viral changes, and which biomarkers provide early indicators of infection or disease progression.”
The knowledge base will serve as an open repository of diagnostic results, along with data regarding the types of diagnostic instruments used to collect samples. It will provide critical insight regarding testing options, as well as new virus strains that are expected to emerge in the months ahead.
“It’s exciting to bring LLNL’s expertise in pathogen surveillance and diagnostics to this important research,” said Kris Kulp, who leads the Laboratory’s Biosciences and Biotechnology Division. “This collaborative research effort leverages DOE investments in LLNL’s biosciences programs, as well as synergistic work at other DOE labs, to provide insight regarding best practices in reliable diagnostic tools and strategies to combat COVID-19.”
- Lisa Valdez
Contact
 Anne M. Stark
Anne M. Stark
[email protected]
(925) 422-9799
Tags
PhysicsPhysical and Life Sciences
Featured Articles