Supercomputers link proteins to drug side effects
LIVERMORE, California - New medications created by pharmaceutical companies have helped millions of Americans alleviate pain and suffering from their medical conditions. However, the drug creation process often misses many side effects that kill at least 100,000 patients a year, according to the journal Nature.
Lawrence Livermore National Laboratory researchers have discovered a high-tech method of using supercomputers to identify proteins that cause medications to have certain adverse drug reactions (ADR) or side effects. They are using high-performance computers (HPC) to process proteins and drug compounds in an algorithm that produces reliable data outside of a laboratory setting for drug discovery.
The team recently published its findings in the journal PLOS ONE, titled "Adverse Drug Reaction Prediction Using Scores Produced by Large-Scale Drug-Protein Target Docking on High-Performance Computer Machines."
"We need to do something to identify these side effects earlier in the drug development cycle to save lives and reduce costs," said Monte LaBute, a researcher from LLNL's Computational Engineering Division and the paper's lead author.
It takes pharmaceutical companies roughly 15 years to bring a new drug to the market, at an average cost of $2 billion. A new drug compound entering Phase I (early stage) testing is estimated to have an 8 percent chance of reaching the market, according to the Food and Drug Administration (FDA).
A typical drug discovery process begins with identifying which proteins are associated with a specific disease. Candidate drug compounds are combined with target proteins in a process known as binding to determine the drug's effectiveness (efficacy) and/or harmful side effects (toxicity). Target proteins are proteins known to bind with drug compounds in order for the pharmaceutical to work.
While this method is able to identify side effects with many target proteins, there are myriad unknown "off-target" proteins that may bind to the candidate drug and could cause unanticipated side effects.
Because it is cost prohibitive to experimentally test a drug candidate against a potentially large set of proteins -- and the list of possible off-targets is not known ahead of time -- pharmaceutical companies usually only test a minimal set of off-target proteins during the early stages of drug discovery. This results in ADRs remaining undetected through the later stages of drug development, such as clinical trials, and possibly making it to the marketplace.
There have been several highly publicized medications with off-target protein side effects that have reached the marketplace. For example, Avandia, an anti-diabetic drug, caused heart attacks in some patients; and Vioxx, an anti-inflammatory medication, caused heart attacks and strokes among certain patient populations. Both therapeutics were recalled because of their side effects.
"There were no indications of side effects of these medications in early testing or clinical trials," LaBute said. "We need a way to determine the safety of such therapeutics before they reach patients. Our work can help direct such drugs to patients who will benefit the most from them with the least amount of side effects."
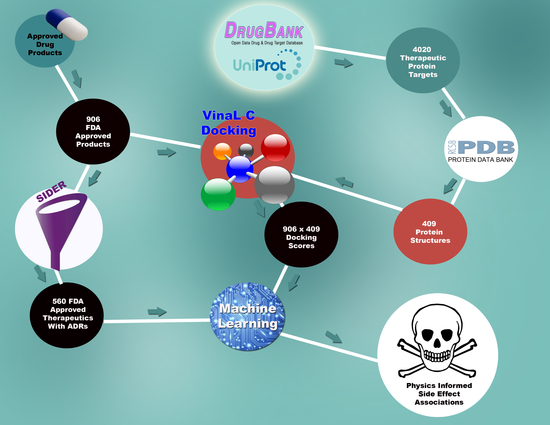
LaBute and the LLNL research team tackled the problem by using supercomputers and information from public databases of drug compounds and proteins. The latter included protein databases of DrugBank,UniProt and Protein Data Bank (PDB), along with drug databases from the FDA and SIDER, which contain FDA-approved drugs with ADRs.
The team examined 4,020 off-target proteins from DrugBank and UniProt. Those proteins were indexed against the PDB, which whittled the number down to 409 off-proteins that have high-quality 3D crystallographic X-ray diffraction structures essential for analysis in a computational setting.
The 409 off-target proteins were fed into a Livermore HPC software known as VinaLC along with 906 FDA-approved drug compounds. VinaLC used a molecular docking matrix that bound the drugs to the proteins. A score was given to each combination to assess whether effective binding occurred.
The binding scores were fed into another computer program and combined with 560 FDA-approved drugs with known side effects. An algorithm was used to determine which proteins were associated with certain side effects.
The Lab team showed that in two categories of disorders -- vascular disorders and neoplasms -- their computational model of predicting side effects in the early stages of drug discovery using off-target proteins was more predictive than current statistical methods that do not include binding scores.
In addition to LLNL ADR prediction methods performing better than current prediction methods, the team's calculations also predicted new potential side effects. For example, they predicted a connection between a protein normally associated with cancer metastasis to vascular disorders like aneurysms. Their ADR predictions were validated by a thorough review of existing scientific data.
"We have discovered a very viable way to find off-target proteins that are important for side effects," LaBute said. "This approach using HPC and molecular docking to find ADRs never really existed before."
The team's findings provide drug companies with a cost-effective and reliable method to screen for side effects, according to LaBute. Their goal is to expand their computational pharmaceutical research to include more off-target proteins for testing and eventually screen every protein in the body.
"If we can do that, the drugs of tomorrow will have less side effects that can potentially lead to fatalities," Labute said. "Optimistically, we could be a decade away from our ultimate goal. However, we need help from pharmaceutical companies, health care providers and the FDA to provide us with patient and therapeutic data."
The LLNL team also includes Felice Lightstone,Xiaohua Zhang, Jason Lenderman, Brian Bennion and Sergio Wong.
Related Links
Adverse Drug Reaction Prediction Using Scores Produced by Large-Scale Drug-Protein Target Docking on High-Performance...PLOS ONE
U.S. Food and Drug Administration
Computational Engineering Division
SIDER 2: Side Effect Resource
DrugBank
UniProt
Protein Data Bank
Tags
HPC, Simulation, and Data ScienceSupercomputer
Computing
Physical and Life Sciences
Featured Articles