LAB REPORT
Science and Technology Making Headlines
April 19, 2019

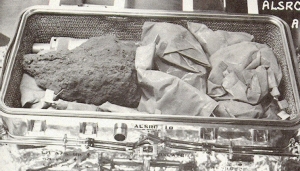
One of the Apollo 16 sample boxes is opened in the Lunar Receiving Laboratory on Earth. The box contains a large rock and many small sample bags. Image courtesy of NASA/Johnson Space Center
Opening up a lunar treasure trove
Armed with new tools and capabilities, NASA has dug some lunar samples out of cold storage to give them a new look. And Lawrence Livermore cosmochemists will be the first ones to take a look.
Nearly 50 years ago, between 1971 and 1972, materials were brought back from the moon to Earth by Apollo astronauts. Nine “special samples” were collected during the Apollo 15, 16 and 17 missions and stored in containers with indium knife-edge seals to maintain a lunar-like vacuum. Apollo mission planners devised these special sample containers to meticulously preserve fragile and transitory sample characteristics (e.g., solar wind volatiles and volatile coatings). Three of these samples have remained sealed in their original Apollo containers until today.
LLNL scientists will get a chance to analyze these Apollo 17 relics to study the geologic history of the site where the rocks were collected, a geologic cold trap where water may have been able to freeze. This marks the first time a sample will be studied in detail since the end of the Apollo program.


Lawrence Livermore scientists have discovered they can use machine learning to automate microencapsulation quality control in real-time, devising an algorithm to determine “good” capsules from “bad” and developing a valve-based mechanism that can sort them without human intervention. Illustration by Jacob Long/LLNL
In real time
Microencapsulated CO2 sorbents (MECS) — tiny, reusable capsules full of a sodium carbonate solution that can absorb carbon dioxide from the air — are a promising technology for capturing carbon from the atmosphere.
To create the caviar-like objects, scientists run three fluids through a series of microfluidic components to create drops that turn into capsules when exposed to ultraviolet light downstream. However, fluid properties and flow rates can change during experiments. These changes can lead to capsules that are defective, improperly sized or otherwise unusable, resulting in device clogging, contaminated samples and wasted time.
To date, this process of creating microcapsules has required constant monitoring, a mundane task for operators. But Lawrence Livermore National Laboratory scientists have discovered they can use machine learning to automate microencapsulation quality control in real-time, devising an algorithm to determine "good" capsules from "bad" and developing a valve-based mechanism that can sort them without human intervention.

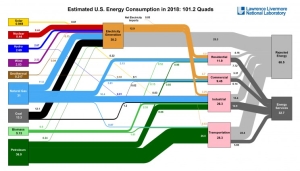
Americans used more energy in 2018 than in any other year, according to the most recent energy flow charts released by Lawrence Livermore National Laboratory.
Energy goes with the flow
Americans used more energy in 2018 than in any other year, according to the most recent energy flow charts released by Lawrence Livermore. Overall total energy consumption rose to 101.2 quadrillion BTU (or "quads"). The prior record, set in 2007, was 101.0 quads. Energy use went up by 3.6 percent from 2017, which also is the largest annual increase since 2010.
Each year, the Laboratory releases energy flow charts that illustrate the nation's consumption and use of energy. Americans used 3.5 quads (quadrillion BTU) more in 2018 than in 2017. A BTU, or British Thermal Unit, is a unit of measurement for energy; 3,400 BTUs is equivalent to about 1 kilowatt-hour.
The largest increases in energy supply came from natural gas, wind and solar energy. In 2018, wind use was up 0.18 quads (7.6 percent) and solar was up 0.18 quads (22 percent). Over the last decade (between 2008 and 2018), total renewable energy production has doubled, including a five-fold increase in wind power and a 48-fold increase in solar. Wind and solar combined now produce more electricity than hydroelectric power, which dominated renewable energy for decades.


A joint LLNL/Sandia National Laboratories team commissioned a Line VISAR diagnostic at Z machine, Sandia’s pulsed power facility. Photo by Randy Montoya/Sandia National Laboratories
The lean, green Z machine
If you’re chasing the elusive goal of nuclear fusion and think you need a bigger reactor to do the job, you first might want to know precisely how much input energy emerging from the wall plug is making it to the heart of your machine.
To better determine energy leaks at Sandia’s powerful Z machine — where remarkable gains in fusion outputs have occurred over the last two and a half decades, including a tripling of output in 2018 — a joint team from Sandia and Lawrence Livermore national laboratories have installed an upgraded laser diagnostic system.
The quest to accurately understand how much power makes it into Z’s fusion reaction has become more pressing as Z moves into producing the huge number of neutrons that now are only a factor of 40 below the milestone where energy output equals energy input, a desirable state known as scientific break-even. The Z machine’s exceptionally large currents — about 26 megamperes — directly compress fusion fuel to the extreme conditions needed for fusion reactions to occur.

Lawrence Livermore National Laboratory has been honored with a Glassdoor Employees’ Choice Award, recognizing it as one of the Best Places to Work in 2019, as well as the top government contractor and top lab.
It’s a great place to work
With historically low unemployment, employers across the nation are aiming to recruit and retain top talent to help their businesses grow. Some employers, of course, are doing a better job than others. Now in its 11th edition, Glassdoor’s 2019 “Best Places to Work” list is based on employee-submitted reviews collected between Oct. 23, 2017 and Oct. 21, 2018. To be eligible, an organization needed to receive at least 75 employee reviews within the past year and must employ 1,000 or more workers.
Employee rankings are based on a 5-point scale ranging from very dissatisfied (1.0) to very satisfied (5.0). Actual calculations extend beyond the thousandth decimal place.
Lawrence Livermore came in with a ranking of 4.4 and placed 24 on the 100 best places to work. “They do not run you into the ground,” writes one reviewer. “They allow you to explore where you're happiest and may pay for additional education needed.” Employer-verified perks include charitable gift matching, tuition assistance and professional development programs.





