The need to rapidly develop new medical countermeasures is critical for enabling a timely response to newly emerging pandemic pathogens, such as COVID-19, and equally for intentional releases of advanced or emerging biological threat agents. LLNL’s efforts in identifying treatment options complement the efforts of the National Institutes of Health, the Department of Defense and other Federal agencies by helping to understand the scientific phenomena contributing to disease and, by extension, to apply these capabilities to help the nation with early-stage medical countermeasure discovery.
Several ongoing research projects have been re-directed or started to support rapid identification of medical countermeasures (drugs and vaccines), including:
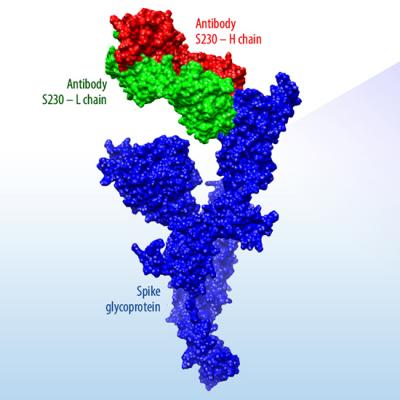
Vaccine and antibody therapeutic optimization pipeline
- The Laboratory has developed an artificial intelligence–driven computational design platform, which is being used to optimize binding of antibodies to the COVID-19 virus. An initial set of promising antibody designs has been identified and is undergoing experimental validation. This platform can also be used to rapidly design vaccine antigens. The initial set of antibody predictions are available on the LLNL COVID-19 Data Portal.
- LLNL researchers have authored an accelerated publication: Rapid in silico design of antibodies targeting SARS-CoV-2 using machine learning and supercomputing
Small molecule (drug) discovery modeling
- The Laboratory is computationally screening millions of commercially available small molecule compounds to identify the subset that can be developed into antiviral drugs for COVID-19. When potential candidates are identified, we conduct additional machine learning screens for improving safety and pharmacokinetic profiles of the potential drugs. Initial small molecule predictions are available on the LLNL COVID-19 Data Portal. Additional information about this effort is available on the LLNL Computing website: Small Molecules, Big Possibilities and Iteration and Scalability.
Structural protein modeling
- We are developing computational drug target identification tools to accelerate the process of finding new drugs. Our initial structural predictions for the SARS-CoV2 spike protein are available on the LLNL COVID-19 Data Portal. Additional information about the data is available on the LLNL Computing website: Open to the Research Community.